| |
| TARGET | MODEL DATA | TEMPLATE |
|---|
Model
Icon | Model/Fold
Reliability | Sequence
Database
Link | Database Annotation | Organism | Protein
Size | Modeled
Segment | Size | Seq
Id(%) | PDB
code | PDB
Segment | PDB Comment |
|---|
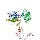 |  | P06715.1 | gshr_ecoli recname: full=glutathione
reductase; short=gr; short=grase | Escherichia coli,
Escherichia coli K-12,
etc. | 450 | 2-450 | 449 | 13 | 1h7wA | 184-624 | dihydropyrimidine dehydrogenase
(dpd) from pig |
 |  | P06715.1 | gshr_ecoli recname: full=glutathione
reductase; short=gr; short=grase | Escherichia coli,
Escherichia coli K-12,
etc. | 450 | 133-303 | 171 | 26 | 3oc4A | 103-277 | |
 |  | P06715.1 | gshr_ecoli recname: full=glutathione
reductase; short=gr; short=grase | Escherichia coli,
Escherichia coli K-12,
etc. | 450 | 2-450 | 449 | 34 | 1h6vA | 10-483 | mammalian thioredoxin
reductase |
 |  | P06715.1 | gshr_ecoli recname: full=glutathione
reductase; short=gr; short=grase | Escherichia coli,
Escherichia coli K-12,
etc. | 450 | 3-450 | 448 | 98 | 1gesA | 3-450 | anatomy of an engineered
nad-binding site |
 |  | 809425 | b chain b, anatomy of an
engineered nad-binding site | Escherichia coli | 450 | 2-450 | 449 | 12 | 1h7wA | 184-624 | dihydropyrimidine dehydrogenase
(dpd) from pig |
 |  | 809425 | b chain b, anatomy of an
engineered nad-binding site | Escherichia coli | 450 | 1-423 | 423 | 10 | 1vqwA | 3-443 | crystal structure of
a protein with similarity
to flavin-containing
monooxygenases and to
mammalian dimethylalanine
monooxygenases |
 |  | P06715.1 | gshr_ecoli recname: full=glutathione
reductase; short=gr; short=grase | Escherichia coli,
Escherichia coli K-12,
etc. | 450 | 3-450 | 448 | 98 | 1gesA | 3-450 | anatomy of an engineered
nad-binding site |
 |  | 809425 | b chain b, anatomy of an
engineered nad-binding site | Escherichia coli | 450 | 1-366 | 366 | 12 | 1djnA | 386-728 | structural and biochemical
characterization of
recombinant wild type
trimethylamine dehydrogenase
from methylophilus methylotrophus
(sp. w3a1) |
 |  | P06715.1 | gshr_ecoli recname: full=glutathione
reductase; short=gr; short=grase | Escherichia coli,
Escherichia coli K-12,
etc. | 450 | 5-444 | 440 | 24 | 3l8kA | 3-446 | crystal structure of
a dihydrolipoyl dehydrogenase
from sulfolobus solfataricus |
 |  | P06715.1 | gshr_ecoli recname: full=glutathione
reductase; short=gr; short=grase | Escherichia coli,
Escherichia coli K-12,
etc. | 450 | 78-449 | 372 | 22 | 3kd9A | 26-433 | crystal structure of
pyridine nucleotide
disulfide oxidoreductase
from pyrococcus horikoshii |
 |  | P06715.1 | gshr_ecoli recname: full=glutathione
reductase; short=gr; short=grase | Escherichia coli,
Escherichia coli K-12,
etc. | 450 | 2-444 | 443 | 21 | 3l8kA | 0-446 | crystal structure of
a dihydrolipoyl dehydrogenase
from sulfolobus solfataricus |
 |  | P06715.1 | gshr_ecoli recname: full=glutathione
reductase; short=gr; short=grase | Escherichia coli,
Escherichia coli K-12,
etc. | 450 | 90-438 | 349 | 23 | 3iwaA | 28-437 | crystal structure of
a fad-dependent pyridine
nucleotide-disulphide
oxidoreductase from
desulfovibrio vulgaris |
 |  | P06715.1 | gshr_ecoli recname: full=glutathione
reductase; short=gr; short=grase | Escherichia coli,
Escherichia coli K-12,
etc. | 450 | 3-450 | 448 | 54 | 3grs | 19-478 | refined structure of
glutathione reductase
at 1.54 angstroms resolution |
 |  | P06715.1 | gshr_ecoli recname: full=glutathione
reductase; short=gr; short=grase | Escherichia coli,
Escherichia coli K-12,
etc. | 450 | 102-442 | 341 | 19 | 3iwaA | 82-449 | crystal structure of
a fad-dependent pyridine
nucleotide-disulphide
oxidoreductase from
desulfovibrio vulgaris |
 |  | P06715.1 | gshr_ecoli recname: full=glutathione
reductase; short=gr; short=grase | Escherichia coli,
Escherichia coli K-12,
etc. | 450 | 1-362 | 362 | 12 | 2gv8A | 3-383 | crystal structure of
flavin-containing monooxygenase
(fmo) from s.pombe and
nadph cofactor complex |
 |  | P06715.1 | gshr_ecoli recname: full=glutathione
reductase; short=gr; short=grase | Escherichia coli,
Escherichia coli K-12,
etc. | 450 | 1-366 | 366 | 13 | 1djnA | 386-728 | structural and biochemical
characterization of
recombinant wild type
trimethylamine dehydrogenase
from methylophilus methylotrophus
(sp. w3a1) |
 |  | P06715.1 | gshr_ecoli recname: full=glutathione
reductase; short=gr; short=grase | Escherichia coli,
Escherichia coli K-12,
etc. | 450 | 1-405 | 405 | 12 | 1vqwA | 3-443 | crystal structure of
a protein with similarity
to flavin-containing
monooxygenases and to
mammalian dimethylalanine
monooxygenases |
 |  | P06715.1 | gshr_ecoli recname: full=glutathione
reductase; short=gr; short=grase | Escherichia coli,
Escherichia coli K-12,
etc. | 450 | 1-450 | 450 | 50 | 3grs | 18-478 | refined structure of
glutathione reductase
at 1.54 angstroms resolution |
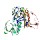 |  | P06715.1 | gshr_ecoli recname: full=glutathione
reductase; short=gr; short=grase | Escherichia coli,
Escherichia coli K-12,
etc. | 450 | 74-446 | 373 | 16 | 3oc4A | 29-426 | |
 |  | P06715.1 | gshr_ecoli recname: full=glutathione
reductase; short=gr; short=grase | Escherichia coli,
Escherichia coli K-12,
etc. | 450 | 3-450 | 448 | 98 | 1gesA | 3-450 | anatomy of an engineered
nad-binding site |
 |  | P06715.1 | gshr_ecoli recname: full=glutathione
reductase; short=gr; short=grase | Escherichia coli,
Escherichia coli K-12,
etc. | 450 | 1-423 | 423 | 10 | 1vqwA | 3-443 | crystal structure of
a protein with similarity
to flavin-containing
monooxygenases and to
mammalian dimethylalanine
monooxygenases |
 |  | P06715.1 | gshr_ecoli recname: full=glutathione
reductase; short=gr; short=grase | Escherichia coli,
Escherichia coli K-12,
etc. | 450 | 1-450 | 450 | 39 | 2jk6A | 1-472 | structure of trypanothione
reductase from leishmania
infantum |
 |  | P06715.1 | gshr_ecoli recname: full=glutathione
reductase; short=gr; short=grase | Escherichia coli,
Escherichia coli K-12,
etc. | 450 | 78-443 | 366 | 26 | 3kd9A | 26-427 | crystal structure of
pyridine nucleotide
disulfide oxidoreductase
from pyrococcus horikoshii |
 |  | P06715.1 | gshr_ecoli recname: full=glutathione
reductase; short=gr; short=grase | Escherichia coli,
Escherichia coli K-12,
etc. | 450 | 1-320 | 320 | 15 | 2gqfA | 1-393 | crystal structure of
hypothetical flavoprotein
hi0933 from haemophilus
influenzae rd. |