| |
| TARGET | MODEL DATA | TEMPLATE |
|---|
Model
Icon | Model/Fold
Reliability | Sequence
Database
Link | Database Annotation | Organism | Protein
Size | Modeled
Segment | Size | Seq
Id(%) | PDB
code | PDB
Segment | PDB Comment |
|---|
 |  | P06959 | recname: full=dihydrolipoyllysine-residue
acetyltransferase component
of pyruvat e dehydrogenase
complex;ec=2.3.1.12;altname:
full=dihydrolipoamide acetyltransferase
component of pyruvate dehydrogenase | Escherichia coli,
Escherichia fergusonii,
etc. | 630 | 398-630 | 233 | 32 | 1sczA | 173-402 | improved structural
model for the catalytic
domain of e.coli dihydrolipoamide
succinyltransferase |
 |  | P06959 | dihydrolipoyllysine-residue
acetyltransferase component
of pyruvatede dehydrog
enase complex (ec 2.3.1.12)
(e2) (dihydrolipoamidede
acetyltransferase component
of pyruvate dehydrogenase
complex) | Escherichia coli | 629 | 199-298 | 100 | 20 | 1fyc | 3-105 | inner lipoyl domain
from human pyruvate
dehydrogenase (pdh)
complex, nmr, 1 structure |
 |  | P06959 | recname: full=dihydrolipoyllysine-residue
acetyltransferase component
of pyruvat e dehydrogenase
complex;ec=2.3.1.12;altname:
full=dihydrolipoamide acetyltransferase
component of pyruvate dehydrogenase | Escherichia coli,
Escherichia fergusonii,
etc. | 630 | 329-364 | 36 | 58 | 2eq8C | 131-166 | crystal structure of
lipoamide dehydrogenase
from thermus thermophilus
hb8 with psbdp |
 |  | P06959 | recname: full=dihydrolipoyllysine-residue
acetyltransferase component
of pyruvat e dehydrogenase
complex;ec=2.3.1.12;altname:
full=dihydrolipoamide acetyltransferase
component of pyruvate dehydrogenase | Escherichia coli,
Escherichia fergusonii,
etc. | 630 | 387-630 | 244 | 55 | 1dpbA | 396-637 | crystallographic and
enzymatic investigations
on the role of ser558,
his610 and asn614 in
the catalytic mechanism
of azotobacter vinelandii
dihydrolipoamide acetyltransferase
(e2p) |
 |  | P06959 | recname: full=dihydrolipoyllysine-residue
acetyltransferase component
of pyruvat e dehydrogenase
complex;ec=2.3.1.12;altname:
full=dihydrolipoamide acetyltransferase
component of pyruvate dehydrogenase | Escherichia coli,
Escherichia fergusonii,
etc. | 630 | 13-79 | 67 | 25 | 2b8fA | 7-73 | solution structure of
bacillus subtilis blap
apo form (energy minimized
mean structure) |
 |  | P06959 | recname: full=dihydrolipoyllysine-residue
acetyltransferase component
of pyruvat e dehydrogenase
complex;ec=2.3.1.12;altname:
full=dihydrolipoamide acetyltransferase
component of pyruvate dehydrogenase | Escherichia coli,
Escherichia fergusonii,
etc. | 630 | 375-630 | 256 | 39 | 1b5sA | 187-425 | dihydrolipoyl transacetylase
(e.c.2.3.1.12) catalytic
domain (residues 184-425)
from bacillus stearothermophilus |
 |  | P06959 | recname: full=dihydrolipoyllysine-residue
acetyltransferase component
of pyruvat e dehydrogenase
complex;ec=2.3.1.12;altname:
full=dihydrolipoamide acetyltransferase
component of pyruvate dehydrogenase | Escherichia coli,
Escherichia fergusonii,
etc. | 630 | 206-293 | 88 | 100 | 2k7vA | 2-89 | deletions in a surface
loop divert the folding
of a protein domain
into a metastable dimeric
form |
 |  | P06959 | recname: full=dihydrolipoyllysine-residue
acetyltransferase component
of pyruvat e dehydrogenase
complex;ec=2.3.1.12;altname:
full=dihydrolipoamide acetyltransferase
component of pyruvate dehydrogenase | Escherichia coli,
Escherichia fergusonii,
etc. | 630 | 372-630 | 259 | 40 | 1b5sA | 184-425 | dihydrolipoyl transacetylase
(e.c.2.3.1.12) catalytic
domain (residues 184-425)
from bacillus stearothermophilus |
 |  | P06959 | dihydrolipoyllysine-residue
acetyltransferase component
of pyruvatede dehydrog
enase complex (ec 2.3.1.12)
(e2) (dihydrolipoamidede
acetyltransferase component
of pyruvate dehydrogenase
complex) | Escherichia coli | 629 | 328-365 | 38 | 47 | 1ebdC | 131-168 | dihydrolipoamide dehydrogenase
complexed with the binding
domain of the dihydrolipoamide
acetylase |
 |  | P06959 | dihydrolipoyllysine-residue
acetyltransferase component
of pyruvatede dehydrog
enase complex (ec 2.3.1.12)
(e2) (dihydrolipoamidede
acetyltransferase component
of pyruvate dehydrogenase
complex) | Escherichia coli | 629 | 385-629 | 245 | 55 | 1eaf | 395-637 | atomic structure of
the cubic core of the
pyruvate dehydrogenase
multienzyme complex |
 |  | P06959 | recname: full=dihydrolipoyllysine-residue
acetyltransferase component
of pyruvat e dehydrogenase
complex;ec=2.3.1.12;altname:
full=dihydrolipoamide acetyltransferase
component of pyruvate dehydrogenase | Escherichia coli,
Escherichia fergusonii,
etc. | 630 | 17-75 | 59 | 32 | 3u9sA | 657-715 | |
 |  | P06959 | recname: full=dihydrolipoyllysine-residue
acetyltransferase component
of pyruvat e dehydrogenase
complex;ec=2.3.1.12;altname:
full=dihydrolipoamide acetyltransferase
component of pyruvate dehydrogenase | Escherichia coli,
Escherichia fergusonii,
etc. | 630 | 205-284 | 80 | 99 | 1qjoA | 1-80 | innermost lipoyl domain
of the pyruvate dehydrogenase
from escherichia coli |
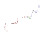 |  | P06959 | recname: full=dihydrolipoyllysine-residue
acetyltransferase component
of pyruvat e dehydrogenase
complex;ec=2.3.1.12;altname:
full=dihydrolipoamide acetyltransferase
component of pyruvate dehydrogenase | Escherichia coli,
Escherichia fergusonii,
etc. | 630 | 206-293 | 88 | 100 | 2k7vA | 2-89 | deletions in a surface
loop divert the folding
of a protein domain
into a metastable dimeric
form |
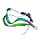 |  | P06959 | recname: full=dihydrolipoyllysine-residue
acetyltransferase component
of pyruvat e dehydrogenase
complex;ec=2.3.1.12;altname:
full=dihydrolipoamide acetyltransferase
component of pyruvate dehydrogenase | Escherichia coli,
Escherichia fergusonii,
etc. | 630 | 16-68 | 53 | 42 | 2d5dA | 87-139 | structure of biotin
carboxyl carrier protein
(74val start) from pyrococcus
horikoshi ot3 ligand
free form ii |
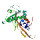 |  | P06959 | recname: full=dihydrolipoyllysine-residue
acetyltransferase component
of pyruvat e dehydrogenase
complex;ec=2.3.1.12;altname:
full=dihydrolipoamide acetyltransferase
component of pyruvate dehydrogenase | Escherichia coli,
Escherichia fergusonii,
etc. | 630 | 424-630 | 207 | 35 | 2ii3A | 213-418 | crystal structure of
a cubic core of the
dihydrolipoamide acyltransferase
(e2b) component in the
branched-chain alpha-ketoacid
dehydrogenase complex
(bckdc), oxidized coenzyme
a-bound form |
 |  | P06959 | recname: full=dihydrolipoyllysine-residue
acetyltransferase component
of pyruvat e dehydrogenase
complex;ec=2.3.1.12;altname:
full=dihydrolipoamide acetyltransferase
component of pyruvate dehydrogenase | Escherichia coli,
Escherichia fergusonii,
etc. | 630 | 45-151 | 107 | 15 | 2k32A | 3-109 | truncated acra from
campylobacter jejuni
for glycosylation studies |
 |  | P06959 | recname: full=dihydrolipoyllysine-residue
acetyltransferase component
of pyruvat e dehydrogenase
complex;ec=2.3.1.12;altname:
full=dihydrolipoamide acetyltransferase
component of pyruvate dehydrogenase | Escherichia coli,
Escherichia fergusonii,
etc. | 630 | 384-629 | 246 | 100 | 4n72A | 383-628 | |
 |  | P06959 | recname: full=dihydrolipoyllysine-residue
acetyltransferase component
of pyruvat e dehydrogenase
complex;ec=2.3.1.12;altname:
full=dihydrolipoamide acetyltransferase
component of pyruvate dehydrogenase | Escherichia coli,
Escherichia fergusonii,
etc. | 630 | 1-50 | 50 | 78 | 2k7vA | 1-50 | deletions in a surface
loop divert the folding
of a protein domain
into a metastable dimeric
form |
 |  | P06959 | recname: full=dihydrolipoyllysine-residue
acetyltransferase component
of pyruvat e dehydrogenase
complex;ec=2.3.1.12;altname:
full=dihydrolipoamide acetyltransferase
component of pyruvate dehydrogenase | Escherichia coli,
Escherichia fergusonii,
etc. | 630 | 119-176 | 58 | 40 | 2d5dA | 87-144 | structure of biotin
carboxyl carrier protein
(74val start) from pyrococcus
horikoshi ot3 ligand
free form ii |
 |  | P06959 | recname: full=dihydrolipoyllysine-residue
acetyltransferase component
of pyruvat e dehydrogenase
complex;ec=2.3.1.12;altname:
full=dihydrolipoamide acetyltransferase
component of pyruvate dehydrogenase | Escherichia coli,
Escherichia fergusonii,
etc. | 630 | 203-282 | 80 | 35 | 1k8mA | 1-82 | solution structure of
the lipoic acid-bearing
domain of the e2 component
of human, mitochondrial
branched-chain alpha-ketoacid
dehydrogenase |
 |  | P06959 | recname: full=dihydrolipoyllysine-residue
acetyltransferase component
of pyruvat e dehydrogenase
complex;ec=2.3.1.12;altname:
full=dihydrolipoamide acetyltransferase
component of pyruvate dehydrogenase | Escherichia coli,
Escherichia fergusonii,
etc. | 630 | 5-80 | 76 | 55 | 1iyuA | 4-78 | lipoyl domain of pyruvate
dehydrogenase complex,
nmr, minimized average
structure |
 |  | P06959 | dihydrolipoyllysine-residue
acetyltransferase component
of pyruvatede dehydrog
enase complex (ec 2.3.1.12)
(e2) (dihydrolipoamidede
acetyltransferase component
of pyruvate dehydrogenase
complex) | Escherichia coli | 629 | 387-629 | 243 | 39 | 1b5sA | 186-425 | dihydrolipoyl transacetylase
(e.c.2.3.1.12) catalytic
domain (residues 184-425)
from bacillus stearothermophilus |
 |  | P06959 | recname: full=dihydrolipoyllysine-residue
acetyltransferase component
of pyruvat e dehydrogenase
complex;ec=2.3.1.12;altname:
full=dihydrolipoamide acetyltransferase
component of pyruvate dehydrogenase | Escherichia coli,
Escherichia fergusonii,
etc. | 630 | 3-79 | 77 | 49 | 1gjxA | 3-80 | solution structure of
the lipoyl domain of
the chimeric dihydrolipoyl
dehydrogenase p64k from
neisseria meningitidis |
 |  | P06959 | dihydrolipoyllysine-residue
acetyltransferase component
of pyruvatede dehydrog
enase complex (ec 2.3.1.12)
(e2) (dihydrolipoamidede
acetyltransferase component
of pyruvate dehydrogenase
complex) | Escherichia coli | 629 | 102-184 | 83 | 29 | 1k8mA | 2-86 | solution structure of
the lipoic acid-bearing
domain of the e2 component
of human, mitochondrial
branched-chain alpha-ketoacid
dehydrogenase |
 |  | P06959 | recname: full=dihydrolipoyllysine-residue
acetyltransferase component
of pyruvat e dehydrogenase
complex;ec=2.3.1.12;altname:
full=dihydrolipoamide acetyltransferase
component of pyruvate dehydrogenase | Escherichia coli,
Escherichia fergusonii,
etc. | 630 | 327-372 | 46 | 100 | 4qoyE | 122-167 | |
 |  | P06959 | dihydrolipoyllysine-residue
acetyltransferase component
of pyruvatede dehydrog
enase complex (ec 2.3.1.12)
(e2) (dihydrolipoamidede
acetyltransferase component
of pyruvate dehydrogenase
complex) | Escherichia coli | 629 | 200-311 | 112 | 16 | 1htp | 17-130 | refined structures at
2 angstroms and 2.2
angstroms of the two
forms of the h-protein,
a lipoamide-containing
protein of the glycine
decarboxylase complex |
 |  | P06959 | dihydrolipoyllysine-residue
acetyltransferase component
of pyruvatede dehydrog
enase complex (ec 2.3.1.12)
(e2) (dihydrolipoamidede
acetyltransferase component
of pyruvate dehydrogenase
complex) | Escherichia coli | 629 | 104-182 | 79 | 85 | 1qjoA | 2-80 | innermost lipoyl domain
of the pyruvate dehydrogenase
from escherichia coli |
 |  | P06959 | recname: full=dihydrolipoyllysine-residue
acetyltransferase component
of pyruvat e dehydrogenase
complex;ec=2.3.1.12;altname:
full=dihydrolipoamide acetyltransferase
component of pyruvate dehydrogenase | Escherichia coli,
Escherichia fergusonii,
etc. | 630 | 386-630 | 245 | 54 | 1dpbA | 395-637 | crystallographic and
enzymatic investigations
on the role of ser558,
his610 and asn614 in
the catalytic mechanism
of azotobacter vinelandii
dihydrolipoamide acetyltransferase
(e2p) |
 |  | P06959 | recname: full=dihydrolipoyllysine-residue
acetyltransferase component
of pyruvat e dehydrogenase
complex;ec=2.3.1.12;altname:
full=dihydrolipoamide acetyltransferase
component of pyruvate dehydrogenase | Escherichia coli,
Escherichia fergusonii,
etc. | 630 | 326-366 | 41 | 46 | 1w85I | 128-168 | the crystal structure
of pyruvate deydrogenase
e1 bound to the peripheral
subunit binding domain
of e2 |
 |  | P06959 | recname: full=dihydrolipoyllysine-residue
acetyltransferase component
of pyruvat e dehydrogenase
complex;ec=2.3.1.12;altname:
full=dihydrolipoamide acetyltransferase
component of pyruvate dehydrogenase | Escherichia coli,
Escherichia fergusonii,
etc. | 630 | 206-293 | 88 | 100 | 2k7vA | 2-89 | deletions in a surface
loop divert the folding
of a protein domain
into a metastable dimeric
form |
 |  | P06959 | recname: full=dihydrolipoyllysine-residue
acetyltransferase component
of pyruvat e dehydrogenase
complex;ec=2.3.1.12;altname:
full=dihydrolipoamide acetyltransferase
component of pyruvate dehydrogenase | Escherichia coli,
Escherichia fergusonii,
etc. | 630 | 386-630 | 245 | 55 | 1eaf | 395-637 | atomic structure of
the cubic core of the
pyruvate dehydrogenase
multienzyme complex |
 |  | P06959 | dihydrolipoyllysine-residue
acetyltransferase component
of pyruvatede dehydrog
enase complex (ec 2.3.1.12)
(e2) (dihydrolipoamidede
acetyltransferase component
of pyruvate dehydrogenase
complex) | Escherichia coli | 629 | 1-73 | 73 | 33 | 1dczA | 54-122 | biotin carboxyl carrier
domain of transcarboxylase
(tc 1.3s) |
 |  | P06959 | recname: full=dihydrolipoyllysine-residue
acetyltransferase component
of pyruvat e dehydrogenase
complex;ec=2.3.1.12;altname:
full=dihydrolipoamide acetyltransferase
component of pyruvate dehydrogenase | Escherichia coli,
Escherichia fergusonii,
etc. | 630 | 105-183 | 79 | 50 | 1iyuA | 1-78 | lipoyl domain of pyruvate
dehydrogenase complex,
nmr, minimized average
structure |
 |  | P06959 | dihydrolipoyllysine-residue
acetyltransferase component
of pyruvatede dehydrog
enase complex (ec 2.3.1.12)
(e2) (dihydrolipoamidede
acetyltransferase component
of pyruvate dehydrogenase
complex) | Escherichia coli | 629 | 204-283 | 80 | 55 | 1gjxA | 1-81 | solution structure of
the lipoyl domain of
the chimeric dihydrolipoyl
dehydrogenase p64k from
neisseria meningitidis |