| |
| TARGET | MODEL DATA | TEMPLATE |
|---|
Model
Icon | Model/Fold
Reliability | Sequence
Database
Link | Database Annotation | Organism | Protein
Size | Modeled
Segment | Size | Seq
Id(%) | PDB
code | PDB
Segment | PDB Comment |
|---|
 |  | A0A086TYR8 | recname: full=1-(5-phosphoribosyl)-5-[(5-phosphoribosylamino)methylideneamino]
i midazole-4-carboxamide
isomerase {eco:0000256|hamap-rule:mf_01014,
eco:0000256|rulebase:ru003658};ec=5.3.1.16
{eco:00002 | Escherichia coli,
Escherichia coli K-12,
etc. | 245 | 2-90 | 89 | 60 | 4gj1A | 4-92 | |
 |  | 41713 | unnamed protein product | Escherichia coli | 245 | 30-245 | 216 | 13 | 1rpxA | 24-226 | d-ribulose-5-phosphate
3-epimerase from solanum
tuberosum chloroplasts |
 |  | 41713 | unnamed protein product | Escherichia coli | 245 | 191-229 | 39 | 21 | 1wv2A | 1182-1220 | crystal structure of
thiamine biosynthesis
protein from pseudomonas
aeruginosa |
 |  | A0A086TYR8 | recname: full=1-(5-phosphoribosyl)-5-[(5-phosphoribosylamino)methylideneamino]
i midazole-4-carboxamide
isomerase {eco:0000256|hamap-rule:mf_01014,
eco:0000256|rulebase:ru003658};ec=5.3.1.16
{eco:00002 | Escherichia coli,
Escherichia coli K-12,
etc. | 245 | 30-245 | 216 | 13 | 1rpxA | 24-226 | d-ribulose-5-phosphate
3-epimerase from solanum
tuberosum chloroplasts |
 |  | 41713 | unnamed protein product | Escherichia coli | 245 | 30-238 | 209 | 13 | 1dvjB | 10-202 | crystal structure of
orotidine monophosphate
decarboxylase complexed
with 6-azaump |
 |  | 41713 | unnamed protein product | Escherichia coli | 245 | 2-244 | 243 | 49 | 4gj1A | 4-243 | |
 |  | A0A086TYR8 | recname: full=1-(5-phosphoribosyl)-5-[(5-phosphoribosylamino)methylideneamino]
i midazole-4-carboxamide
isomerase {eco:0000256|hamap-rule:mf_01014,
eco:0000256|rulebase:ru003658};ec=5.3.1.16
{eco:00002 | Escherichia coli,
Escherichia coli K-12,
etc. | 245 | 2-244 | 243 | 28 | 1ka9F | 7-244 | imidazole glycerol phosphate
synthase |
 |  | A0A086TYR8 | recname: full=1-(5-phosphoribosyl)-5-[(5-phosphoribosylamino)methylideneamino]
i midazole-4-carboxamide
isomerase {eco:0000256|hamap-rule:mf_01014,
eco:0000256|rulebase:ru003658};ec=5.3.1.16
{eco:00002 | Escherichia coli,
Escherichia coli K-12,
etc. | 245 | 191-229 | 39 | 21 | 1wv2A | 1182-1220 | crystal structure of
thiamine biosynthesis
protein from pseudomonas
aeruginosa |
 |  | A0A086TYR8 | recname: full=1-(5-phosphoribosyl)-5-[(5-phosphoribosylamino)methylideneamino]
i midazole-4-carboxamide
isomerase {eco:0000256|hamap-rule:mf_01014,
eco:0000256|rulebase:ru003658};ec=5.3.1.16
{eco:00002 | Escherichia coli,
Escherichia coli K-12,
etc. | 245 | 30-238 | 209 | 13 | 1gox | 107-312 | refined structure of
spinach glycolate oxidase
at 2 angstroms resolution |
 |  | A0A086TYR8 | recname: full=1-(5-phosphoribosyl)-5-[(5-phosphoribosylamino)methylideneamino]
i midazole-4-carboxamide
isomerase {eco:0000256|hamap-rule:mf_01014,
eco:0000256|rulebase:ru003658};ec=5.3.1.16
{eco:00002 | Escherichia coli,
Escherichia coli K-12,
etc. | 245 | 2-223 | 222 | 32 | 1h5yA | 7-225 | hisf protein from pyrobaculum
aerophilum |
 |  | A0A086TYR8 | recname: full=1-(5-phosphoribosyl)-5-[(5-phosphoribosylamino)methylideneamino]
i midazole-4-carboxamide
isomerase {eco:0000256|hamap-rule:mf_01014,
eco:0000256|rulebase:ru003658};ec=5.3.1.16
{eco:00002 | Escherichia coli,
Escherichia coli K-12,
etc. | 245 | 74-240 | 167 | 17 | 1eepA | 104-260 | 2.4 a resolution crystal
structure of borrelia
burgdorferi inosine
5'-monphosphate dehydrogenase
in complex with a sulfate
ion |
 |  | A0A086TYR8 | recname: full=1-(5-phosphoribosyl)-5-[(5-phosphoribosylamino)methylideneamino]
i midazole-4-carboxamide
isomerase {eco:0000256|hamap-rule:mf_01014,
eco:0000256|rulebase:ru003658};ec=5.3.1.16
{eco:00002 | Escherichia coli,
Escherichia coli K-12,
etc. | 245 | 2-245 | 244 | 25 | 1h5yA | 7-247 | hisf protein from pyrobaculum
aerophilum |
 |  | A0A086TYR8 | recname: full=1-(5-phosphoribosyl)-5-[(5-phosphoribosylamino)methylideneamino]
i midazole-4-carboxamide
isomerase {eco:0000256|hamap-rule:mf_01014,
eco:0000256|rulebase:ru003658};ec=5.3.1.16
{eco:00002 | Escherichia coli,
Escherichia coli K-12,
etc. | 245 | 1-242 | 242 | 12 | 1kbiA | 202-454 | crystallographic study
of the recombinant flavin-binding
domain of baker's yeast
flavocytochrome b2:
comparison with the
intact wild-type enzyme |
 |  | A0A086TYR8 | recname: full=1-(5-phosphoribosyl)-5-[(5-phosphoribosylamino)methylideneamino]
i midazole-4-carboxamide
isomerase {eco:0000256|hamap-rule:mf_01014,
eco:0000256|rulebase:ru003658};ec=5.3.1.16
{eco:00002 | Escherichia coli,
Escherichia coli K-12,
etc. | 245 | 30-240 | 211 | 16 | 1eunA | 28-198 | structure of 2-keto-3-deoxy-6-phosphogluconate
aldolase from escherichia
coli |
 |  | A0A086TYR8 | recname: full=1-(5-phosphoribosyl)-5-[(5-phosphoribosylamino)methylideneamino]
i midazole-4-carboxamide
isomerase {eco:0000256|hamap-rule:mf_01014,
eco:0000256|rulebase:ru003658};ec=5.3.1.16
{eco:00002 | Escherichia coli,
Escherichia coli K-12,
etc. | 245 | 1-244 | 244 | 32 | 2y88A | 5-245 | |
 |  | A0A086TYR8 | recname: full=1-(5-phosphoribosyl)-5-[(5-phosphoribosylamino)methylideneamino]
i midazole-4-carboxamide
isomerase {eco:0000256|hamap-rule:mf_01014,
eco:0000256|rulebase:ru003658};ec=5.3.1.16
{eco:00002 | Escherichia coli,
Escherichia coli K-12,
etc. | 245 | 1-240 | 240 | 10 | 1to3A | 16-267 | structure of yiht from
salmonella typhimurium |
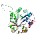 |  | A0A086TYR8 | recname: full=1-(5-phosphoribosyl)-5-[(5-phosphoribosylamino)methylideneamino]
i midazole-4-carboxamide
isomerase {eco:0000256|hamap-rule:mf_01014,
eco:0000256|rulebase:ru003658};ec=5.3.1.16
{eco:00002 | Escherichia coli,
Escherichia coli K-12,
etc. | 245 | 2-241 | 240 | 12 | 1pii | 39-252 | three-dimensional structure
of the bifunctional
enzyme phosphoribosylanthranilate
isomerase: indoleglycerolphosphate
synthase from escherichia
coli refined at 2.0
angstroms resolution |
 |  | A0A086TYR8 | recname: full=1-(5-phosphoribosyl)-5-[(5-phosphoribosylamino)methylideneamino]
i midazole-4-carboxamide
isomerase {eco:0000256|hamap-rule:mf_01014,
eco:0000256|rulebase:ru003658};ec=5.3.1.16
{eco:00002 | Escherichia coli,
Escherichia coli K-12,
etc. | 245 | 1-245 | 245 | 14 | 1b3oA | 36-463 | ternary complex of human
type-ii inosine monophosphate
dehydrogenase with 6-cl-imp
and selenazole adenine
dinucleotide |
 |  | 41713 | unnamed protein product | Escherichia coli | 245 | 2-240 | 239 | 10 | 1ak5 | 36-261 | inosine monophosphate
dehydrogenase (impdh)
from tritrichomonas
foetus |
 |  | 41713 | unnamed protein product | Escherichia coli | 245 | 2-129 | 128 | 16 | 1ep3A | 171-297 | crystal structure of
lactococcus lactis dihydroorotate
dehydrogenase b. data
collected under cryogenic
conditions. |
 |  | 41713 | unnamed protein product | Escherichia coli | 245 | 30-238 | 209 | 13 | 1gox | 107-312 | refined structure of
spinach glycolate oxidase
at 2 angstroms resolution |
 |  | A0A086TYR8 | recname: full=1-(5-phosphoribosyl)-5-[(5-phosphoribosylamino)methylideneamino]
i midazole-4-carboxamide
isomerase {eco:0000256|hamap-rule:mf_01014,
eco:0000256|rulebase:ru003658};ec=5.3.1.16
{eco:00002 | Escherichia coli,
Escherichia coli K-12,
etc. | 245 | 2-244 | 243 | 50 | 4gj1A | 4-243 | |
 |  | A0A086TYR8 | recname: full=1-(5-phosphoribosyl)-5-[(5-phosphoribosylamino)methylideneamino]
i midazole-4-carboxamide
isomerase {eco:0000256|hamap-rule:mf_01014,
eco:0000256|rulebase:ru003658};ec=5.3.1.16
{eco:00002 | Escherichia coli,
Escherichia coli K-12,
etc. | 245 | 2-240 | 239 | 24 | 1thfD | 6-239 | cyclase subunit of imidazoleglycerolphosphate
synthase from thermotoga
maritima |
 |  | 41713 | unnamed protein product | Escherichia coli | 245 | 2-240 | 239 | 24 | 1thfD | 6-239 | cyclase subunit of imidazoleglycerolphosphate
synthase from thermotoga
maritima |
 |  | A0A086TYR8 | recname: full=1-(5-phosphoribosyl)-5-[(5-phosphoribosylamino)methylideneamino]
i midazole-4-carboxamide
isomerase {eco:0000256|hamap-rule:mf_01014,
eco:0000256|rulebase:ru003658};ec=5.3.1.16
{eco:00002 | Escherichia coli,
Escherichia coli K-12,
etc. | 245 | 2-129 | 128 | 16 | 1ep3A | 171-297 | crystal structure of
lactococcus lactis dihydroorotate
dehydrogenase b. data
collected under cryogenic
conditions. |
 |  | A0A086TYR8 | recname: full=1-(5-phosphoribosyl)-5-[(5-phosphoribosylamino)methylideneamino]
i midazole-4-carboxamide
isomerase {eco:0000256|hamap-rule:mf_01014,
eco:0000256|rulebase:ru003658};ec=5.3.1.16
{eco:00002 | Escherichia coli,
Escherichia coli K-12,
etc. | 245 | 1-240 | 240 | 27 | 1qo2A | 2-238 | crystal structure of
n-((5'-phosphoribosyl)-formimino)-5-aminoimidazol-4-carboxamid
ribonucleotid isomerase
(ec 3.1.3.15., hisa) |
 |  | A0A086TYR8 | recname: full=1-(5-phosphoribosyl)-5-[(5-phosphoribosylamino)methylideneamino]
i midazole-4-carboxamide
isomerase {eco:0000256|hamap-rule:mf_01014,
eco:0000256|rulebase:ru003658};ec=5.3.1.16
{eco:00002 | Escherichia coli,
Escherichia coli K-12,
etc. | 245 | 29-113 | 85 | 19 | 1wv2A | 1143-1227 | crystal structure of
thiamine biosynthesis
protein from pseudomonas
aeruginosa |
 |  | 41713 | unnamed protein product | Escherichia coli | 245 | 74-240 | 167 | 17 | 1eepA | 104-260 | 2.4 a resolution crystal
structure of borrelia
burgdorferi inosine
5'-monphosphate dehydrogenase
in complex with a sulfate
ion |
 |  | A0A086TYR8 | recname: full=1-(5-phosphoribosyl)-5-[(5-phosphoribosylamino)methylideneamino]
i midazole-4-carboxamide
isomerase {eco:0000256|hamap-rule:mf_01014,
eco:0000256|rulebase:ru003658};ec=5.3.1.16
{eco:00002 | Escherichia coli,
Escherichia coli K-12,
etc. | 245 | 30-238 | 209 | 17 | 2tpsA | 31-212 | thiamin phosphate synthase |
 |  | 41713 | unnamed protein product | Escherichia coli | 245 | 30-240 | 211 | 16 | 1eunA | 28-198 | structure of 2-keto-3-deoxy-6-phosphogluconate
aldolase from escherichia
coli |
 |  | A0A086TYR8 | recname: full=1-(5-phosphoribosyl)-5-[(5-phosphoribosylamino)methylideneamino]
i midazole-4-carboxamide
isomerase {eco:0000256|hamap-rule:mf_01014,
eco:0000256|rulebase:ru003658};ec=5.3.1.16
{eco:00002 | Escherichia coli,
Escherichia coli K-12,
etc. | 245 | 1-244 | 244 | 93 | 5a5wA | 1-244 | |
 |  | 41713 | unnamed protein product | Escherichia coli | 245 | 1-240 | 240 | 27 | 1qo2A | 2-238 | crystal structure of
n-((5'-phosphoribosyl)-formimino)-5-aminoimidazol-4-carboxamid
ribonucleotid isomerase
(ec 3.1.3.15., hisa) |
 |  | A0A086TYR8 | recname: full=1-(5-phosphoribosyl)-5-[(5-phosphoribosylamino)methylideneamino]
i midazole-4-carboxamide
isomerase {eco:0000256|hamap-rule:mf_01014,
eco:0000256|rulebase:ru003658};ec=5.3.1.16
{eco:00002 | Escherichia coli,
Escherichia coli K-12,
etc. | 245 | 2-240 | 239 | 13 | 1a53 | 36-246 | complex of indole-3-glycerolphosphate
synthase from sulfolobus
solfataricus with indole-3-glycerolphosphate
at 2.0 a resolution |
 |  | 41713 | unnamed protein product | Escherichia coli | 245 | 2-240 | 239 | 13 | 1a53 | 36-246 | complex of indole-3-glycerolphosphate
synthase from sulfolobus
solfataricus with indole-3-glycerolphosphate
at 2.0 a resolution |
 |  | A0A086TYR8 | recname: full=1-(5-phosphoribosyl)-5-[(5-phosphoribosylamino)methylideneamino]
i midazole-4-carboxamide
isomerase {eco:0000256|hamap-rule:mf_01014,
eco:0000256|rulebase:ru003658};ec=5.3.1.16
{eco:00002 | Escherichia coli,
Escherichia coli K-12,
etc. | 245 | 2-243 | 242 | 25 | 1thfD | 6-242 | cyclase subunit of imidazoleglycerolphosphate
synthase from thermotoga
maritima |
 |  | 41713 | unnamed protein product | Escherichia coli | 245 | 2-223 | 222 | 32 | 1h5yA | 7-225 | hisf protein from pyrobaculum
aerophilum |
 |  | A0A086TYR8 | recname: full=1-(5-phosphoribosyl)-5-[(5-phosphoribosylamino)methylideneamino]
i midazole-4-carboxamide
isomerase {eco:0000256|hamap-rule:mf_01014,
eco:0000256|rulebase:ru003658};ec=5.3.1.16
{eco:00002 | Escherichia coli,
Escherichia coli K-12,
etc. | 245 | 71-110 | 40 | 42 | 1w0mA | 171-210 | triosephosphate isomerase
from thermoproteus tenax |
 |  | 41713 | unnamed protein product | Escherichia coli | 245 | 30-238 | 209 | 17 | 2tpsA | 31-212 | thiamin phosphate synthase |
 |  | A0A086TYR8 | recname: full=1-(5-phosphoribosyl)-5-[(5-phosphoribosylamino)methylideneamino]
i midazole-4-carboxamide
isomerase {eco:0000256|hamap-rule:mf_01014,
eco:0000256|rulebase:ru003658};ec=5.3.1.16
{eco:00002 | Escherichia coli,
Escherichia coli K-12,
etc. | 245 | 30-238 | 209 | 13 | 1dvjB | 10-202 | crystal structure of
orotidine monophosphate
decarboxylase complexed
with 6-azaump |
 |  | A0A086TYR8 | recname: full=1-(5-phosphoribosyl)-5-[(5-phosphoribosylamino)methylideneamino]
i midazole-4-carboxamide
isomerase {eco:0000256|hamap-rule:mf_01014,
eco:0000256|rulebase:ru003658};ec=5.3.1.16
{eco:00002 | Escherichia coli,
Escherichia coli K-12,
etc. | 245 | 75-229 | 155 | 13 | 3ktsA | 7-181 | crystal structure of
glycerol uptake operon
antiterminator regulatory
protein from listeria
monocytogenes str. 4b
f2365 |
 |  | A0A086TYR8 | recname: full=1-(5-phosphoribosyl)-5-[(5-phosphoribosylamino)methylideneamino]
i midazole-4-carboxamide
isomerase {eco:0000256|hamap-rule:mf_01014,
eco:0000256|rulebase:ru003658};ec=5.3.1.16
{eco:00002 | Escherichia coli,
Escherichia coli K-12,
etc. | 245 | 62-110 | 49 | 20 | 1to3A | 208-260 | structure of yiht from
salmonella typhimurium |
 |  | 41713 | unnamed protein product | Escherichia coli | 245 | 2-241 | 240 | 12 | 1pii | 39-252 | three-dimensional structure
of the bifunctional
enzyme phosphoribosylanthranilate
isomerase: indoleglycerolphosphate
synthase from escherichia
coli refined at 2.0
angstroms resolution |
 |  | 41713 | unnamed protein product | Escherichia coli | 245 | 29-113 | 85 | 20 | 1wv2A | 1143-1227 | crystal structure of
thiamine biosynthesis
protein from pseudomonas
aeruginosa |
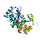 |  | A0A086TYR8 | recname: full=1-(5-phosphoribosyl)-5-[(5-phosphoribosylamino)methylideneamino]
i midazole-4-carboxamide
isomerase {eco:0000256|hamap-rule:mf_01014,
eco:0000256|rulebase:ru003658};ec=5.3.1.16
{eco:00002 | Escherichia coli,
Escherichia coli K-12,
etc. | 245 | 2-245 | 244 | 28 | 1ka9F | 7-245 | imidazole glycerol phosphate
synthase |
 |  | A0A086TYR8 | recname: full=1-(5-phosphoribosyl)-5-[(5-phosphoribosylamino)methylideneamino]
i midazole-4-carboxamide
isomerase {eco:0000256|hamap-rule:mf_01014,
eco:0000256|rulebase:ru003658};ec=5.3.1.16
{eco:00002 | Escherichia coli,
Escherichia coli K-12,
etc. | 245 | 1-240 | 240 | 27 | 1qo2A | 2-238 | crystal structure of
n-((5'-phosphoribosyl)-formimino)-5-aminoimidazol-4-carboxamid
ribonucleotid isomerase
(ec 3.1.3.15., hisa) |