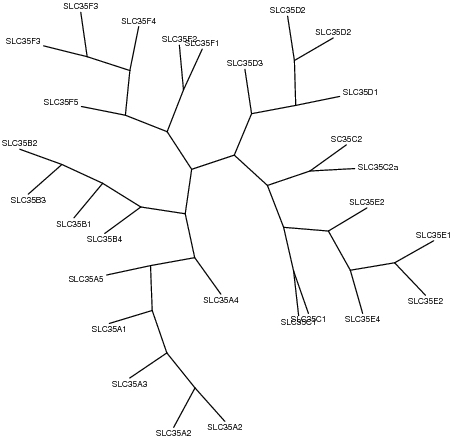
Phylogenetic trees were constructed using PhyML and drawn using Phylip, based on multiple sequence alignments by MUSCLE.
oooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooo
--- PhyML v3.0 (179M) ---
http://www.atgc-montpellier.fr/phyml
Copyright CNRS - Universite Montpellier II
oooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooo
. Sequence filename: slc35-muscle.fa.fas.phylip
. Data set: #1
. Tree topology search : NNIs
. Initial tree: BioNJ
. Model of amino acids substitution: LG
. Number of taxa: 28
. Log-likelihood: -42622.65626
. Unconstrained likelihood: -9481.63602
. Parsimony: 9567
. Tree size: 45.13239
. Discrete gamma model: Yes
- Number of categories: 4
- Gamma shape parameter: 2.498
. Time used 0h12m19s
. 739 seconds
oooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooo
Suggested citation:
S. Guindon & O. Gascuel
"A simple, fast, and accurate algorithm to estimate large phylogenies by maximum likelihood"
Systematic Biology. 2003. 52(5):696-704.
oooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooooo